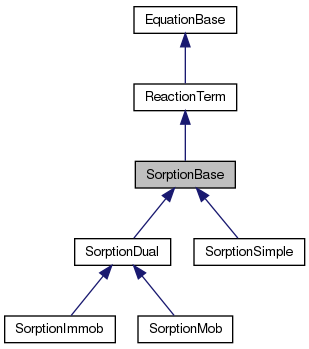
#include <sorption_base.hh>
Classes | |
| class | EqData |
Public Member Functions | |
| SorptionBase (Mesh &init_mesh, Input::Record in_rec) | |
| virtual | ~SorptionBase (void) |
| void | zero_time_step () override |
| virtual void | update_solution (void) |
| void | set_porosity (Field< 3, FieldValue< 3 >::Scalar > &por_m) |
| void | make_tables (void) |
| void | output_data (void) override |
| Output method. More... | |
| void | output_vector_gather (void) override |
 Public Member Functions inherited from ReactionTerm Public Member Functions inherited from ReactionTerm | |
| ReactionTerm (Mesh &init_mesh, Input::Record in_rec) | |
| Constructor. More... | |
| ~ReactionTerm (void) | |
| Destructor. More... | |
| void | choose_next_time (void) override |
| Disable changes in TimeGovernor by empty method. More... | |
| ReactionTerm & | names (const std::vector< string > &names) |
| Sets the names of substances considered in transport. More... | |
| ReactionTerm & | output_stream (OutputTime &ostream) |
| Sets the output stream which is given from transport class. More... | |
| ReactionTerm & | concentration_matrix (double **concentration, Distribution *conc_distr, int *el_4_loc, int *row_4_el) |
 Public Member Functions inherited from EquationBase Public Member Functions inherited from EquationBase | |
| EquationBase () | |
| EquationBase (Mesh &mesh, const Input::Record in_rec) | |
| virtual | ~EquationBase () |
| virtual void | initialize () |
| Initialize fields. More... | |
| virtual void | set_time_upper_constraint (double dt) |
| virtual void | set_time_lower_constraint (double dt) |
| TimeGovernor const & | time () |
| virtual void | set_time_governor (TimeGovernor &time) |
| double | planned_time () |
| double | solved_time () |
| Mesh & | mesh () |
| TimeMark::Type | mark_type () |
| FieldSet & | data () |
| virtual void | get_solution_vector (double *&vector, unsigned int &size) |
| virtual void | get_parallel_solution_vector (Vec &vector) |
Static Public Member Functions | |
| static Input::Type::Selection | make_output_selection (const string &output_field_name, const string &selection_name) |
Static Public Attributes | |
| static Input::Type::Record | input_type |
 Static Public Attributes inherited from ReactionTerm Static Public Attributes inherited from ReactionTerm | |
| static Input::Type::AbstractRecord | input_type = AbstractRecord("ReactionTerm", "Equation for reading information about simple chemical reactions.") |
| static Input::Type::Record | input_type_output_record |
| Specification of the output record. More... | |
Protected Member Functions | |
| SorptionBase () | |
| void | initialize_substance_ids (const std::vector< string > &names, Input::Record in_rec) |
| void | init_from_input (Input::Record in_rec) override |
| Initializes private members of sorption from the input record. More... | |
| void | init_from_input_reaction (Input::Record in_rec) |
| double ** | compute_reaction (double **concentrations, int loc_el) |
| virtual void | isotherm_reinit (std::vector< Isotherm > &isotherms, const ElementAccessor< 3 > &elm)=0 |
| void | print_sorption_parameters (void) |
| void | allocate_output_mpi (void) |
Protected Attributes | |
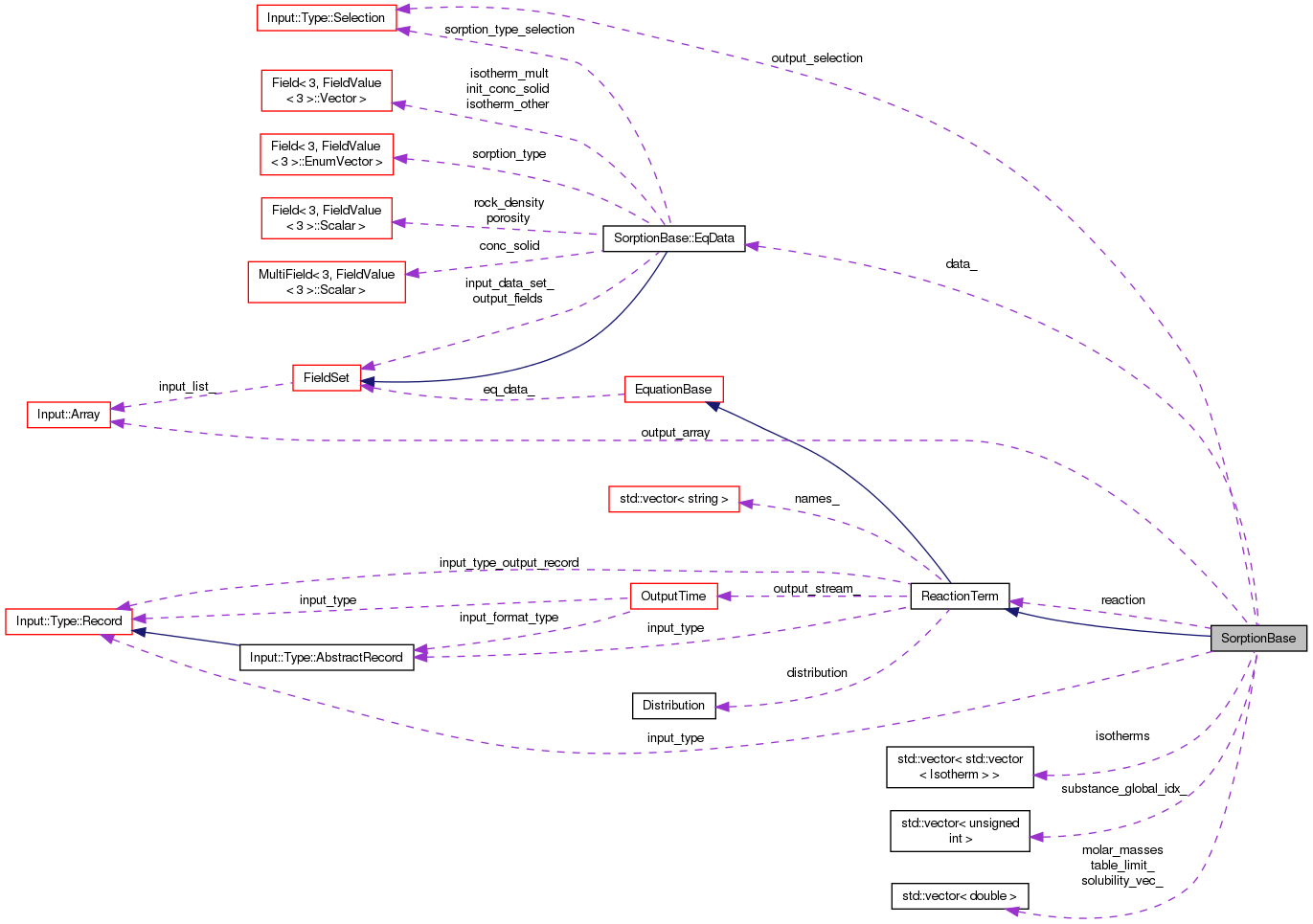
| EqData * | data_ |
| int | nr_of_regions |
| int | nr_of_points |
| std::vector< double > | molar_masses |
| double | solvent_density |
| std::vector< double > | solubility_vec_ |
| std::vector< double > | table_limit_ |
| std::vector< std::vector < Isotherm > > | isotherms |
| unsigned int | n_substances_ |
| std::vector< unsigned int > | substance_global_idx_ |
| Mapping from local indexing of substances to global. More... | |
| double ** | conc_solid |
| Input::Array | output_array |
| Input::Type::Selection | output_selection |
| ReactionTerm * | reaction |
members used in output routines | |
| Vec * | vconc_solid |
| PETSC sorbed concentration vector (parallel). More... | |
| Vec * | vconc_solid_out |
| PETSC sorbed concentration vector output (gathered - sequential) More... | |
| double ** | conc_solid_out |
| sorbed concentration array output (gathered - sequential) More... | |
 Protected Attributes inherited from ReactionTerm Protected Attributes inherited from ReactionTerm | |
| double ** | concentration_matrix_ |
| int * | el_4_loc |
| Indices of elements belonging to local dofs. More... | |
| int * | row_4_el |
| Indices of rows belonging to elements. More... | |
| Distribution * | distribution |
| Pointer to reference to distribution of elements between processors. More... | |
| vector< string > | names_ |
| Names belonging to substances. More... | |
| OutputTime * | output_stream_ |
| Pointer to a transport output stream. More... | |
 Protected Attributes inherited from EquationBase Protected Attributes inherited from EquationBase | |
| bool | equation_empty_ |
| flag is true if only default constructor was called More... | |
| Mesh * | mesh_ |
| TimeGovernor * | time_ |
| Input::Record | input_record_ |
| FieldSet * | eq_data_ |
Detailed Description
Definition at line 22 of file sorption_base.hh.
Constructor & Destructor Documentation
| SorptionBase::SorptionBase | ( | Mesh & | init_mesh, |
| Input::Record | in_rec | ||
| ) |
Constructor with parameter for initialization of a new declared class member
Definition at line 83 of file sorption_base.cc.
|
virtual |
Destructor.
Definition at line 90 of file sorption_base.cc.
|
protected |
Meaningless inherited method. This method disables to use constructor without parameters.
Member Function Documentation
|
protected |
|
protectedvirtual |
For simulation of sorption in just one element either inside of MOBILE or IMMOBILE pores.
Reimplemented from ReactionTerm.
Definition at line 381 of file sorption_base.cc.
|
overrideprotectedvirtual |
Initializes private members of sorption from the input record.
Reimplemented from ReactionTerm.
Definition at line 154 of file sorption_base.cc.

|
protected |
Initializes possible following reactions from input record. It should be called after setting mesh, time_governor, distribution and concentration_matrix if there are some setting methods for reactions called (they are not at the moment, so it could be part of init_from_input).
Definition at line 307 of file sorption_base.cc.
|
protected |
|
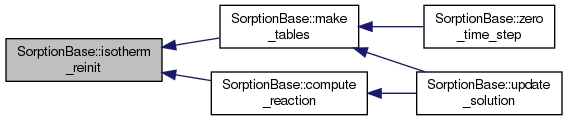
protectedpure virtual |
Implemented in SorptionDual, SorptionSimple, SorptionImmob, and SorptionMob.
|
inlinestatic |
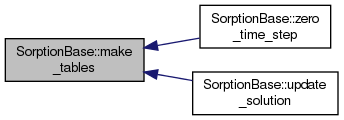
| void SorptionBase::make_tables | ( | void | ) |
Creates interpolation table for isotherms.
Definition at line 361 of file sorption_base.cc.
|
overridevirtual |
Output method.
Some reaction models have their own data to output (sorption, dual porosity) - this is where it must be solved. On the other hand, some do not have (linear reaction, pade approximant) - that is why it is not pure virtual.
Reimplemented from ReactionTerm.
Definition at line 483 of file sorption_base.cc.
|
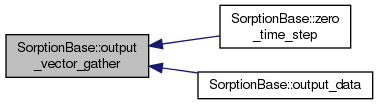
overridevirtual |
Communicate parallel concentration vectors into sequential output vector.
Reimplemented from ReactionTerm.
Definition at line 463 of file sorption_base.cc.
|
protected |
or printing parameters of isotherms under consideration, not necessary to store
Definition at line 429 of file sorption_base.cc.
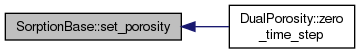
| void SorptionBase::set_porosity | ( | Field< 3, FieldValue< 3 >::Scalar > & | por_m) |
Initialization routines that are done in constructors of descendants. Method data() which access EqData is pure virtual and cannot be called from the base constructor. Sets porosity field - makes a field copy from transport.
Definition at line 423 of file sorption_base.cc.

|
virtual |
Prepared to compute sorption inside all of considered elements. It calls compute_reaction(...) for all the elements controled by concrete processor, when the computation is paralelized.
Reimplemented from EquationBase.
Definition at line 340 of file sorption_base.cc.
|
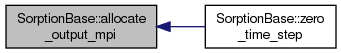
overridevirtual |
Initialization of the solution in the zero time. There is lot of things that can not be done in the constructor since we have not fully initialized fields yet. Fields coming from coupling has to be set after the constructor and before zero_time_step.
Reimplemented from EquationBase.
Definition at line 202 of file sorption_base.cc.
Member Data Documentation
|
protected |
Array for storage infos about sorbed species concentrations.
Definition at line 184 of file sorption_base.hh.
|
protected |
sorbed concentration array output (gathered - sequential)
Definition at line 198 of file sorption_base.hh.
|
protected |
Definition at line 141 of file sorption_base.hh.
|
static |
Static variable for new input data types input
Definition at line 28 of file sorption_base.hh.
|
protected |
Three dimensional array contains intersections between isotherms and mass balance lines. It describes behaviour of sorbents in mobile pores of various rock matrix enviroments. Up to |nr_of_region x nr_of_substances x n_points| doubles. Because of equidistant step lenght in cocidered system of coordinates, just function values are stored.
Definition at line 174 of file sorption_base.hh.
|
protected |
Molar masses of dissolved species (substances)
Definition at line 154 of file sorption_base.hh.
|
protected |
Definition at line 176 of file sorption_base.hh.
|
protected |
Temporary nr_of_points can be computed using step_length. Should be |nr_of_region x nr_of_substances| matrix later.
Definition at line 150 of file sorption_base.hh.
|
protected |
Number of regions.
Definition at line 146 of file sorption_base.hh.
|
protected |
Definition at line 186 of file sorption_base.hh.
|
protected |
Definition at line 188 of file sorption_base.hh.
|
protected |
Reaction model that follows the sorption.
Definition at line 192 of file sorption_base.hh.
|
protected |
Critical concentrations of species dissolved in water.
Definition at line 163 of file sorption_base.hh.
|
protected |
Density of the solvent. TODO: Could be done region dependent, easily.
Definition at line 159 of file sorption_base.hh.
|
protected |
Mapping from local indexing of substances to global.
Definition at line 179 of file sorption_base.hh.
|
protected |
Concentration table limits of species dissolved in water.
Definition at line 167 of file sorption_base.hh.
|
protected |
PETSC sorbed concentration vector (parallel).
Definition at line 196 of file sorption_base.hh.
|
protected |
PETSC sorbed concentration vector output (gathered - sequential)
Definition at line 197 of file sorption_base.hh.
The documentation for this class was generated from the following files:
- /home/jb/workspace/flow123d/src/reaction/sorption_base.hh
- /home/jb/workspace/flow123d/src/reaction/sorption_base.cc
 1.8.4
1.8.4