#include <sorption_base.hh>
Classes | |
| struct | CommonElementData |
| class | EqData |
Public Member Functions | |
| TYPEDEF_ERR_INFO (EI_ArrayName, std::string) | |
| DECLARE_INPUT_EXCEPTION (ExcSubstanceCountMatch,<< "The size of the input array "<< EI_ArrayName::qval<< " does not match the number of substances.") | |
| SorptionBase (Mesh &init_mesh, Input::Record in_rec) | |
| virtual | ~SorptionBase (void) |
| void | initialize () override |
| Prepares the object to usage. More... | |
| void | zero_time_step () override |
| void | update_solution (void) override |
| Updates the solution. More... | |
| void | output_data (void) override |
| Output method. More... | |
| bool | evaluate_time_constraint (double &time_constraint) override |
| Computes a constraint for time step. More... | |
 Public Member Functions inherited from ReactionTerm Public Member Functions inherited from ReactionTerm | |
| TYPEDEF_ERR_INFO (EI_Substance, std::string) | |
| TYPEDEF_ERR_INFO (EI_Model, std::string) | |
| DECLARE_INPUT_EXCEPTION (ExcUnknownSubstance,<< "Unknown substance name: "<< EI_Substance::qval) | |
| DECLARE_INPUT_EXCEPTION (ExcWrongDescendantModel,<< "Impossible descendant model: "<< EI_Model::qval) | |
| ReactionTerm (Mesh &init_mesh, Input::Record in_rec) | |
| Constructor. More... | |
| ~ReactionTerm (void) | |
| Destructor. More... | |
| void | choose_next_time (void) override |
| Disable changes in TimeGovernor by empty method. More... | |
| ReactionTerm & | substances (SubstanceList &substances) |
| Sets the names of substances considered in transport. More... | |
| ReactionTerm & | output_stream (std::shared_ptr< OutputTime > ostream) |
| Sets the output stream which is given from transport class. More... | |
| ReactionTerm & | concentration_matrix (double **concentration, Distribution *conc_distr, LongIdx *el_4_loc, LongIdx *row_4_el) |
 Public Member Functions inherited from EquationBase Public Member Functions inherited from EquationBase | |
| EquationBase () | |
| EquationBase (Mesh &mesh, const Input::Record in_rec) | |
| virtual | ~EquationBase () |
| virtual void | set_time_upper_constraint (double dt, std::string message) |
| virtual void | set_time_lower_constraint (double dt, std::string message) |
| TimeGovernor & | time () |
| virtual void | set_time_governor (TimeGovernor &time) |
| double | planned_time () |
| double | solved_time () |
| Mesh & | mesh () |
| TimeMark::Type | mark_type () |
| FieldSet & | data () |
| virtual void | get_solution_vector (double *&vector, unsigned int &size) |
| virtual void | get_parallel_solution_vector (Vec &vector) |
Static Public Member Functions | |
| static const Input::Type::Record & | get_input_type () |
| static Input::Type::Instance | make_output_type (const string &equation_name, const string &output_field_name) |
 Static Public Member Functions inherited from ReactionTerm Static Public Member Functions inherited from ReactionTerm | |
| static Input::Type::Abstract & | it_abstract_term () |
| static Input::Type::Abstract & | it_abstract_mobile_term () |
| static Input::Type::Abstract & | it_abstract_immobile_term () |
| static Input::Type::Abstract & | it_abstract_reaction () |
Protected Member Functions | |
| SorptionBase () | |
| void | make_reactions () |
| void | initialize_substance_ids () |
| Reads names of substances from input and creates indexing to global vector of substance. More... | |
| void | initialize_from_input () |
| Initializes private members of sorption from the input record. More... | |
| void | initialize_fields () |
| Initializes field sets. More... | |
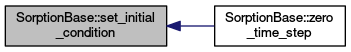
| void | set_initial_condition () |
| Reads and sets initial condition for concentration in solid. More... | |
| void | allocate_output_mpi (void) |
| Allocates petsc vectors, prepares them for output and creates vector scatter. More... | |
| void | output_vector_gather (void) override |
| Gathers all the parallel vectors to enable them to be output. More... | |
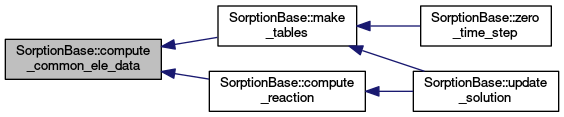
| double ** | compute_reaction (double **concentrations, int loc_el) override |
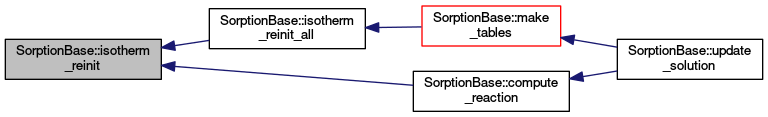
| void | isotherm_reinit (unsigned int i_subst, const ElementAccessor< 3 > &elm) |
| Reinitializes the isotherm. More... | |
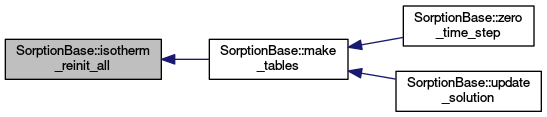
| void | isotherm_reinit_all (const ElementAccessor< 3 > &elm) |
Calls isotherm_reinit for all isotherms. More... | |
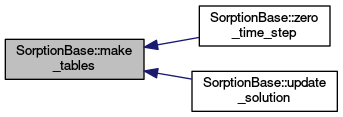
| void | make_tables (void) |
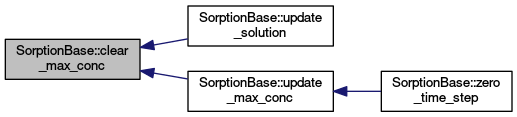
| void | update_max_conc () |
| Computes maximal aqueous concentration at the current step. More... | |
| void | clear_max_conc () |
| Sets max conc to zeros on all regins. More... | |
| virtual void | compute_common_ele_data (const ElementAccessor< 3 > &elem)=0 |
Protected Attributes | |
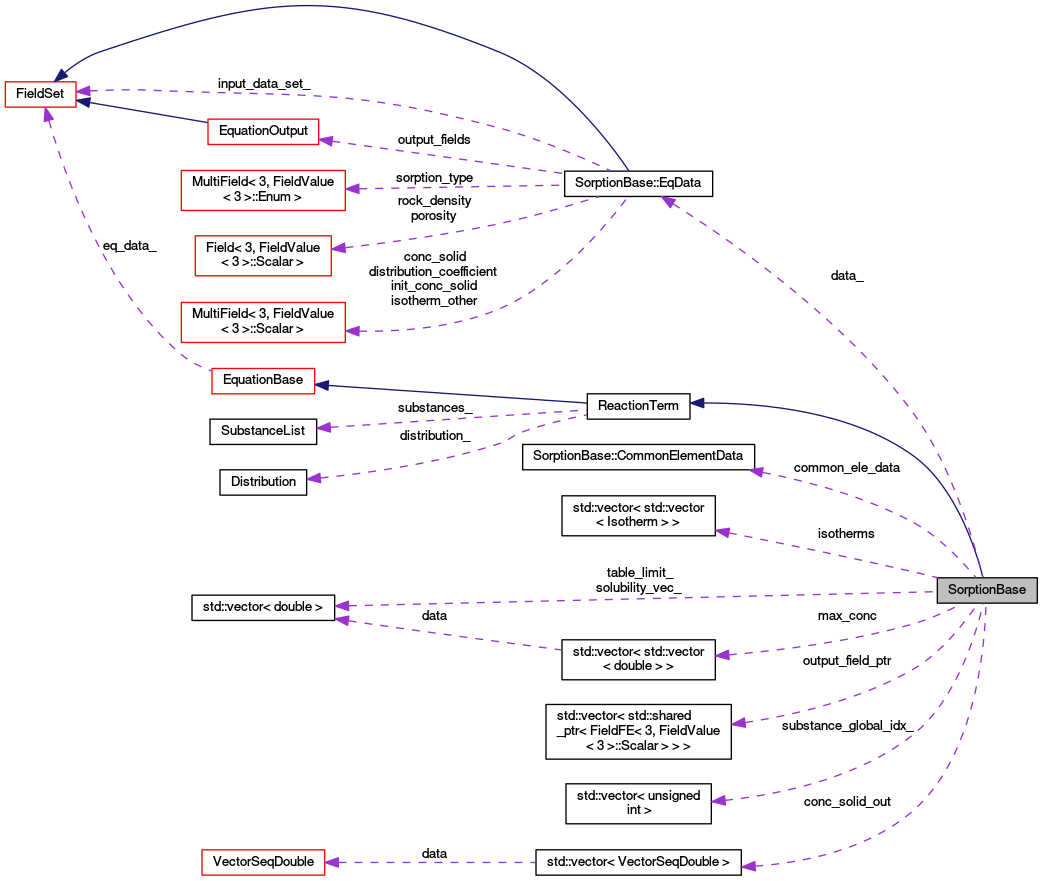
| EqData * | data_ |
| Pointer to equation data. The object is constructed in descendants. More... | |
| unsigned int | n_interpolation_steps_ |
| double | solvent_density_ |
| std::vector< double > | solubility_vec_ |
| std::vector< double > | table_limit_ |
| std::vector< std::vector< double > > | max_conc |
| std::vector< std::vector< Isotherm > > | isotherms |
| unsigned int | n_substances_ |
| std::vector< unsigned int > | substance_global_idx_ |
| Mapping from local indexing of substances to global. More... | |
| double ** | conc_solid |
| std::shared_ptr< ReactionTerm > | reaction_liquid |
| std::shared_ptr< ReactionTerm > | reaction_solid |
| std::vector< std::shared_ptr< FieldFE< 3, FieldValue< 3 >::Scalar > > > | output_field_ptr |
Fields correspond with conc_solid_out. More... | |
| struct SorptionBase::CommonElementData | common_ele_data |
members used in output routines | |
| VecScatter | vconc_out_scatter |
| Output vector scatter. More... | |
| Vec * | vconc_solid |
| PETSC sorbed concentration vector (parallel). More... | |
| std::vector< VectorSeqDouble > | conc_solid_out |
| sorbed concentration array output (gathered - sequential) More... | |
 Protected Attributes inherited from ReactionTerm Protected Attributes inherited from ReactionTerm | |
| double ** | concentration_matrix_ |
| LongIdx * | el_4_loc_ |
| Indices of elements belonging to local dofs. More... | |
| LongIdx * | row_4_el_ |
| Indices of rows belonging to elements. More... | |
| Distribution * | distribution_ |
| Pointer to reference to distribution of elements between processors. More... | |
| SubstanceList | substances_ |
| Names belonging to substances. More... | |
| std::shared_ptr< OutputTime > | output_stream_ |
| Pointer to a transport output stream. More... | |
 Protected Attributes inherited from EquationBase Protected Attributes inherited from EquationBase | |
| bool | equation_empty_ |
| flag is true if only default constructor was called More... | |
| Mesh * | mesh_ |
| TimeGovernor * | time_ |
| Input::Record | input_record_ |
| FieldSet * | eq_data_ |
| std::shared_ptr< Balance > | balance_ |
| object for calculation and writing the mass balance to file. More... | |
Detailed Description
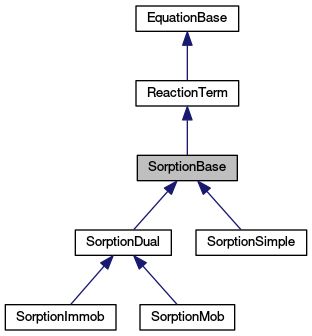
Definition at line 58 of file sorption_base.hh.
Constructor & Destructor Documentation
| SorptionBase::SorptionBase | ( | Mesh & | init_mesh, |
| Input::Record | in_rec | ||
| ) |
Constructor with parameter for initialization of a new declared class member
Definition at line 114 of file sorption_base.cc.
|
virtual |
Destructor.
Definition at line 123 of file sorption_base.cc.
|
protected |
This method disables to use constructor without parameters.
Member Function Documentation
|
protected |
Allocates petsc vectors, prepares them for output and creates vector scatter.
Definition at line 600 of file sorption_base.cc.

|
protected |
Sets max conc to zeros on all regins.
Definition at line 460 of file sorption_base.cc.
|
protectedpure virtual |
Computes CommonElementData. Is pure virtual, implemented differently for simple/mobile/immobile sorption class.
Implemented in SorptionImmob, SorptionMob, SorptionDual, and SorptionSimple.
|
overrideprotectedvirtual |
For simulation of sorption in just one element either inside of MOBILE or IMMOBILE pores.
Implements ReactionTerm.
Definition at line 552 of file sorption_base.cc.

| SorptionBase::DECLARE_INPUT_EXCEPTION | ( | ExcSubstanceCountMatch | , |
| << "The size of the input array "<< EI_ArrayName::qval<< " does not match the number of substances." | |||
| ) |
|
inlineoverridevirtual |
Computes a constraint for time step.
Implements ReactionTerm.
Definition at line 145 of file sorption_base.hh.
|
static |
Static variable for new input data types input
Definition at line 56 of file sorption_base.cc.

|
overridevirtual |
Prepares the object to usage.
Allocating memory, reading input, initialization of fields.
Reimplemented from EquationBase.
Definition at line 169 of file sorption_base.cc.
|
protected |
Initializes field sets.
Definition at line 324 of file sorption_base.cc.

|
protected |
Initializes private members of sorption from the input record.
Definition at line 275 of file sorption_base.cc.

|
protected |
Reads names of substances from input and creates indexing to global vector of substance.
Also creates the local vector of molar masses.
Definition at line 229 of file sorption_base.cc.

|
protected |
Reinitializes the isotherm.
On data change the isotherm is recomputed, possibly new interpolation table is made. NOTE: Be sure to update common element data (porosity, rock density etc.) by compute_common_ele_data(), before calling reinitialization!
Definition at line 422 of file sorption_base.cc.
|
protected |
Calls isotherm_reinit for all isotherms.
Definition at line 452 of file sorption_base.cc.
|
inlinestatic |
|
protected |
Initializes possible following reactions from input record. It should be called after setting mesh, time_governor, distribution and concentration_matrix if there are some setting methods for reactions called (they are not at the moment, so it could be part of init_from_input).
Definition at line 142 of file sorption_base.cc.

|
protected |
Creates interpolation table for isotherms.
Definition at line 487 of file sorption_base.cc.
|
overridevirtual |
Output method.
Some reaction models have their own data to output (sorption, dual porosity)
- this is where it must be reimplemented. On the other hand, some do not have (linear reaction, pade approximant)
- that is why it is not pure virtual.
Reimplemented from ReactionTerm.
Definition at line 634 of file sorption_base.cc.

|
overrideprotectedvirtual |
Gathers all the parallel vectors to enable them to be output.
Reimplemented from ReactionTerm.
Definition at line 623 of file sorption_base.cc.

|
protected |
Reads and sets initial condition for concentration in solid.
Definition at line 385 of file sorption_base.cc.
| SorptionBase::TYPEDEF_ERR_INFO | ( | EI_ArrayName | , |
| std::string | |||
| ) |
|
protected |
Computes maximal aqueous concentration at the current step.
Definition at line 471 of file sorption_base.cc.
|
overridevirtual |
Updates the solution.
Goes through local distribution of elements and calls compute_reaction.
Reimplemented from EquationBase.
Definition at line 402 of file sorption_base.cc.
|
overridevirtual |
Does first computation after initialization process. The time is set and initial condition is set and output.
Reimplemented from EquationBase.
Definition at line 357 of file sorption_base.cc.
Member Data Documentation
|
protected |
|
protected |
Array for storage infos about sorbed species concentrations.
Definition at line 247 of file sorption_base.hh.
|
protected |
sorbed concentration array output (gathered - sequential)
Definition at line 264 of file sorption_base.hh.
|
protected |
Pointer to equation data. The object is constructed in descendants.
Definition at line 207 of file sorption_base.hh.
|
protected |
Three dimensional array contains intersections between isotherms and mass balance lines. It describes behaviour of sorbents in mobile pores of various rock matrix enviroments. Up to |nr_of_region x nr_of_substances x n_points| doubles. Because of equidistant step lenght in cocidered system of coordinates, just function values are stored.
Definition at line 237 of file sorption_base.hh.
|
protected |
Maximum concentration per region. It is used for optimization of interpolation table.
Definition at line 230 of file sorption_base.hh.
|
protected |
Temporary nr_of_points can be computed using step_length. Should be |nr_of_region x nr_of_substances| matrix later.
Definition at line 212 of file sorption_base.hh.
|
protected |
Definition at line 239 of file sorption_base.hh.
|
protected |
Fields correspond with conc_solid_out.
Definition at line 270 of file sorption_base.hh.
|
protected |
Reaction model that follows the sorption.
Definition at line 256 of file sorption_base.hh.
|
protected |
Definition at line 257 of file sorption_base.hh.
|
protected |
Critical concentrations of species dissolved in water.
Definition at line 221 of file sorption_base.hh.
|
protected |
Density of the solvent. TODO: Could be done region dependent, easily.
Definition at line 217 of file sorption_base.hh.
|
protected |
Mapping from local indexing of substances to global.
Definition at line 242 of file sorption_base.hh.
|
protected |
Concentration table limits of species dissolved in water.
Definition at line 225 of file sorption_base.hh.
|
protected |
Output vector scatter.
Definition at line 261 of file sorption_base.hh.
|
protected |
PETSC sorbed concentration vector (parallel).
Definition at line 263 of file sorption_base.hh.
The documentation for this class was generated from the following files:
- /opt/flow123d/flow123d/src/reaction/sorption_base.hh
- /opt/flow123d/flow123d/src/reaction/sorption_base.cc
 1.8.11
1.8.11