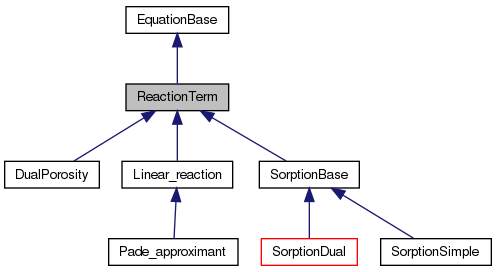
#include <reaction.hh>
Public Member Functions | |
| ReactionTerm (Mesh &init_mesh, Input::Record in_rec) | |
| ~ReactionTerm (void) | |
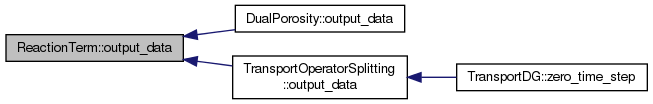
| virtual void | output_data (void) |
Setters | |
| ReactionTerm & | names (const std::vector< string > &names) |
| ReactionTerm & | output_stream (OutputTime &ostream) |
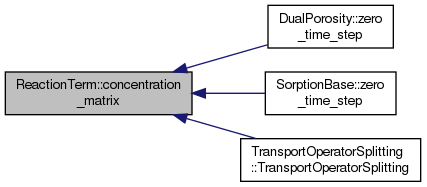
| ReactionTerm & | concentration_matrix (double **concentration, Distribution *conc_distr, int *el_4_loc, int *row_4_el) |
Inherited and not used. | |
TODO: make default empty implementation in EquationBase | |
| virtual void | choose_next_time (void) |
| virtual void | get_parallel_solution_vector (Vec &vc) |
| virtual void | get_solution_vector (double *&vector, unsigned int &size) |
 Public Member Functions inherited from EquationBase Public Member Functions inherited from EquationBase | |
| EquationBase () | |
| EquationBase (Mesh &mesh, const Input::Record in_rec) | |
| virtual | ~EquationBase () |
| virtual void | zero_time_step () |
| virtual void | update_solution () |
| virtual void | set_time_upper_constraint (double dt) |
| virtual void | set_time_lower_constraint (double dt) |
| TimeGovernor const & | time () |
| virtual void | set_time_governor (TimeGovernor &time) |
| double | planned_time () |
| double | solved_time () |
| Mesh & | mesh () |
| TimeMark::Type | mark_type () |
| FieldSet & | data () |
Static Public Attributes | |
| static Input::Type::AbstractRecord | input_type = AbstractRecord("ReactionTerm", "Equation for reading information about simple chemical reactions.") |
| static Input::Type::Record | input_type_output_record |
Protected Member Functions | |
| virtual void | output_vector_gather (void) |
| virtual double ** | compute_reaction (double **concentrations, int loc_el) |
| virtual void | init_from_input (Input::Record in_rec) |
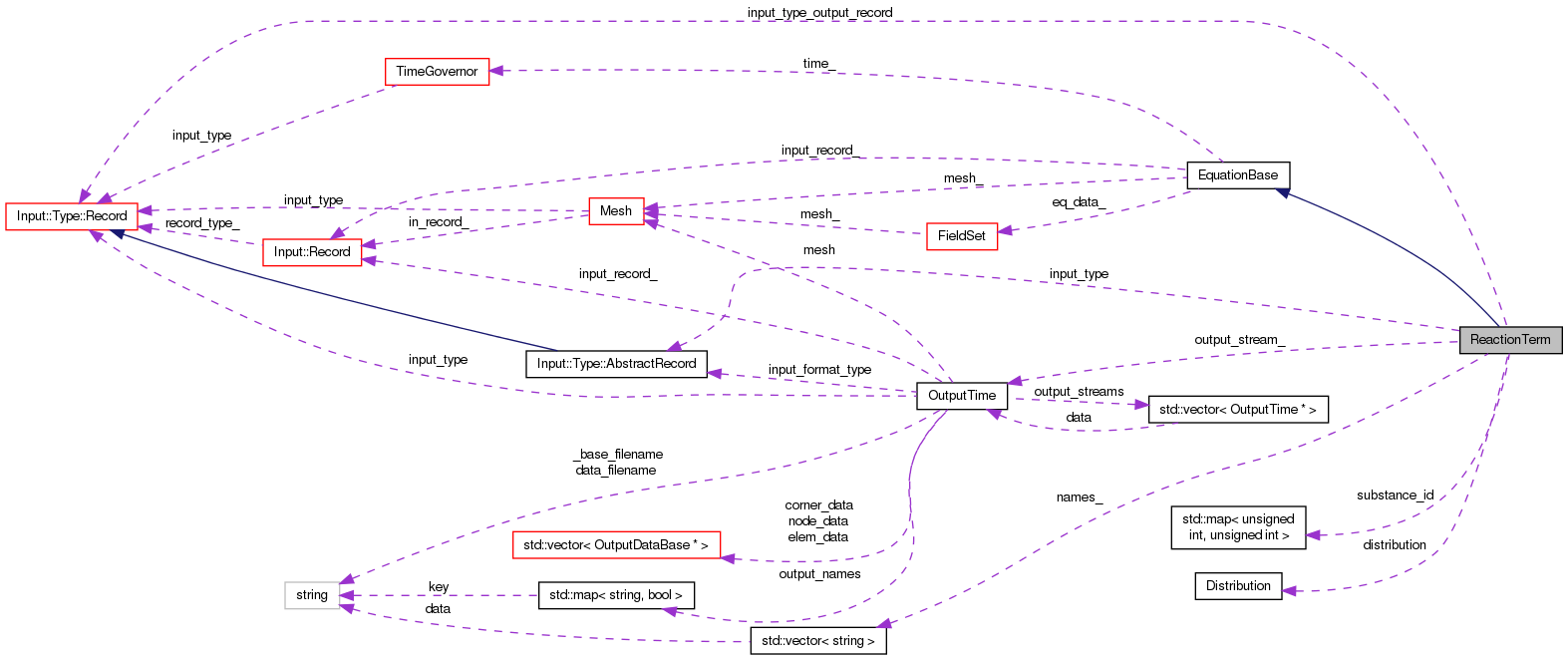
Protected Attributes | |
| double ** | concentration_matrix_ |
| int * | el_4_loc |
| int * | row_4_el |
| Distribution * | distribution |
| vector< string > | names_ |
| std::map< unsigned int, unsigned int > | substance_id |
| Mapping from local indexing of substances to global. More... | |
| OutputTime * | output_stream_ |
| Pointer to a transport output stream. More... | |
 Protected Attributes inherited from EquationBase Protected Attributes inherited from EquationBase | |
| Mesh * | mesh_ |
| TimeGovernor * | time_ |
| Input::Record | input_record_ |
| FieldSet * | eq_data_ |
Detailed Description
Definition at line 21 of file reaction.hh.
Constructor & Destructor Documentation
| ReactionTerm::ReactionTerm | ( | Mesh & | init_mesh, |
| Input::Record | in_rec | ||
| ) |
Constructor with parameter for initialization of a new declared class member TODO: parameter description
Definition at line 24 of file reaction.cc.
| ReactionTerm::~ReactionTerm | ( | void | ) |
Destructor.
Definition at line 35 of file reaction.cc.
Member Function Documentation
|
virtual |
Computation of one time step is split into update_solution() and choose_next_time() in order to allow dependency of the next time step on other coupled models. Fix the next discrete time for computation. Can be rewritten in child class to set possible constrains according to possible equation coefficients or other data which can be result of another model.
Reimplemented from EquationBase.
Definition at line 55 of file reaction.cc.
|
protectedvirtual |
For simulation of chemical reaction in one element only. Inputs should be loc_el and local copies of concentrations of the element, which is then returned.
Reimplemented in SorptionBase, DualPorosity, Pade_approximant, and Linear_reaction.
Definition at line 41 of file reaction.cc.
|
inline |
Sets the concentration matrix for the mobile zone, all substances and on all elements.
Definition at line 59 of file reaction.hh.
|
virtual |
Child class have to implement getter for parallel solution vector. OBSOLETE
Implements EquationBase.
Definition at line 47 of file reaction.cc.
|
virtual |
Child class have to implement getter for sequential solution vector. OBSOLETE
Implements EquationBase.
Definition at line 51 of file reaction.cc.
|
inlineprotectedvirtual |
Initialize data from record in input file. It is intended to use in ascendants.
Reimplemented in SorptionBase, and Linear_reaction.
Definition at line 100 of file reaction.hh.

|
inline |
|
inlinevirtual |
Output method. Some reaction models have their own data to output (sorption, dual porosity) - this is where it must be solved. On the other hand, some do not have (linear reaction, pade approximant) - that is why it is not pure virtual.
Implements EquationBase.
Reimplemented in SorptionBase, and DualPorosity.
Definition at line 73 of file reaction.hh.
|
inline |
|
inlineprotectedvirtual |
Communicate parallel concentration vectors into sequential output vector.
Reimplemented in SorptionBase, and DualPorosity.
Definition at line 89 of file reaction.hh.
Member Data Documentation
|
protected |
Pointer to two-dimensional array[species][elements] containing mobile concentrations.
Definition at line 100 of file reaction.hh.
|
protected |
Pointer to reference to distribution of elements between processors.
Definition at line 119 of file reaction.hh.
|
protected |
Indices of elements belonging to local dofs.
Definition at line 110 of file reaction.hh.
|
static |
Static variable for definition of common input record in reactions.
Definition at line 28 of file reaction.hh.
|
static |
Specification of the output record. Need not to be used by all reaction models, but they should allow output of similar fields.
Definition at line 34 of file reaction.hh.
|
protected |
Names belonging to substances. Should be same as in the transport.
Definition at line 124 of file reaction.hh.
|
protected |
Pointer to a transport output stream.
Definition at line 130 of file reaction.hh.
|
protected |
Indices of rows belonging to elements.
Definition at line 114 of file reaction.hh.
|
protected |
Mapping from local indexing of substances to global.
Definition at line 127 of file reaction.hh.
The documentation for this class was generated from the following files:
- /home/jb/workspace/flow123d/src/reaction/reaction.hh
- /home/jb/workspace/flow123d/src/reaction/reaction.cc
 1.8.4
1.8.4